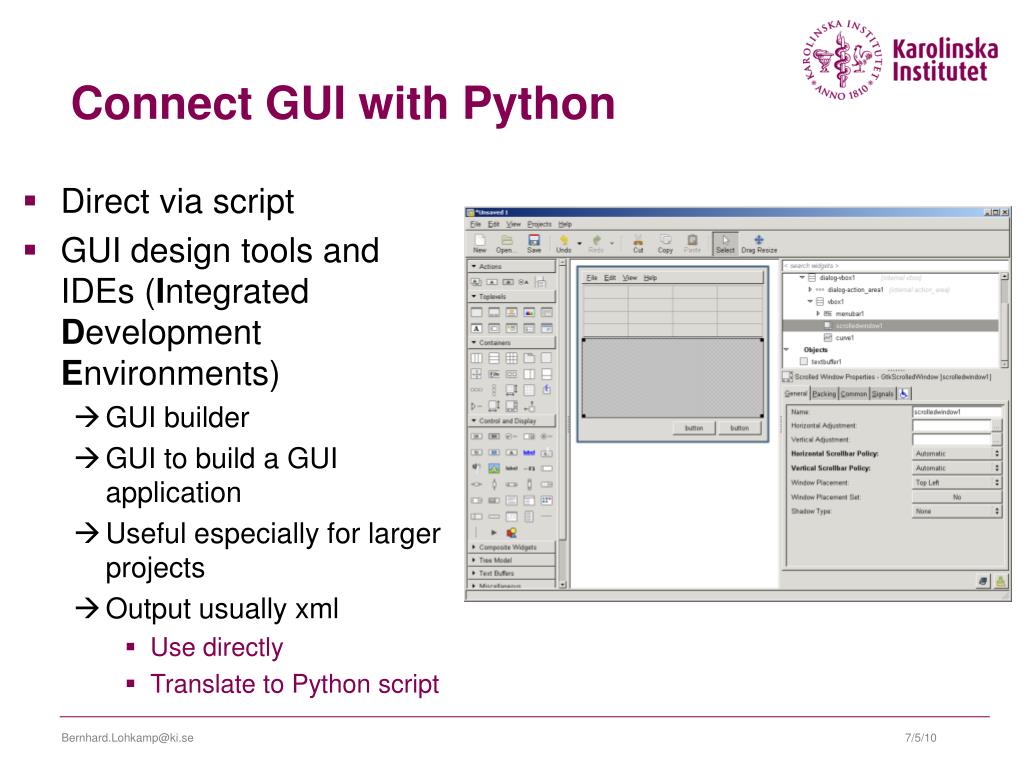
 
A task bar on the right side of the screen allows one to select how to display selected objects.For example, you can save the graphical representation of the molecule into an image file. Pull-down menus provide access to the file functions, the molecular editor, the program options, and a few more advanced tasks.You can interact with the program via four complimentary ways: As of early 2010, development of PyMOL is continued by Schrodinger, LLC. PyMOL is available for many computer platforms either as a pre-compiled binary or in the free source code form.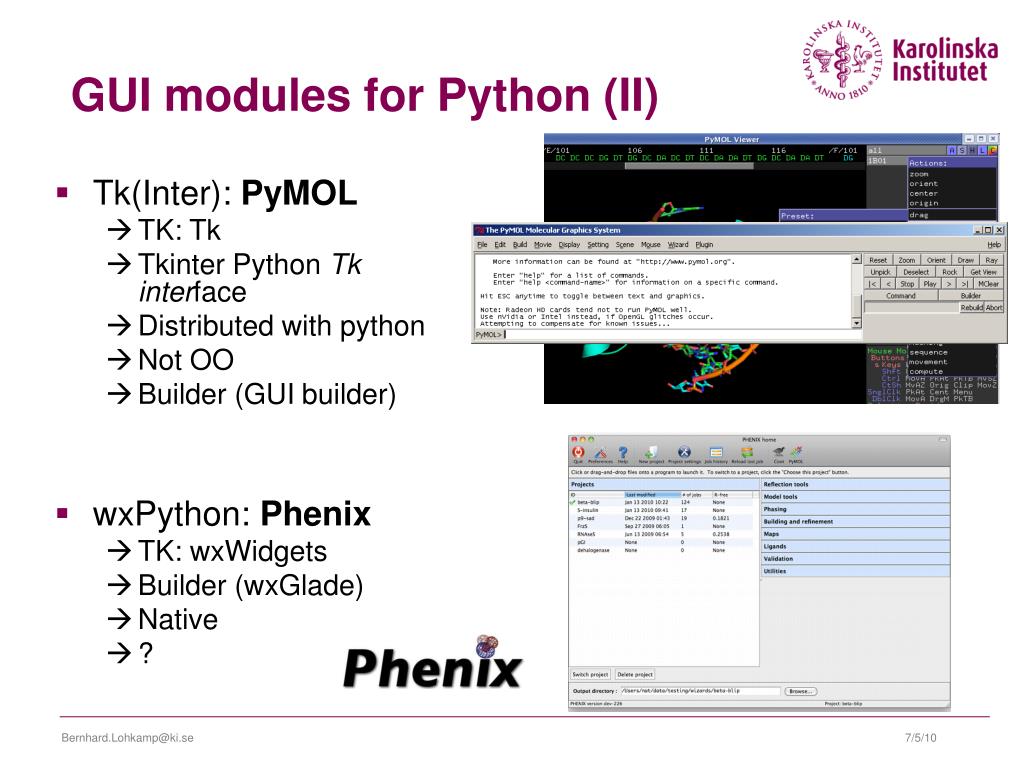 
In more difficult cases you can ask help from PyMOL users via the PyMOL forum. Examples of PyMOL images with commands used to generate these views are available in the Biomolecular Images and Movies for Teaching page. The PyMOL commands are explained in the PyMOL Wiki. As a result, it takes some time to learn the program. 
During the early development, emphasis was placed on providing functionality, and not on intuitive user interface. The strengths of PyMOL arise from the emphasis on high-quality graphics, the use of the powerful programming language ( Python), and the extendability with Plugins. It can also perform many other valuable tasks (such as editing PDB files) that assist you in your research. 
Warren DeLano, PyMOL is a molecular graphics system with an embedded Python interpreter designed for real-time visualization and rapid generation of high-quality molecular graphics images and animations. You will learn about the layout and functionality of the program through a set of guided tasks. This tutorial shows how to effectively use the program PyMOL for visualizing biological macromolecules. Computational analysis of binding modes and protein contacts reported that CD147 and ACE2 might be two complementary receptors mediating virus infection and confirmed the experimental results previously.CSUPERB Tutorial: Molecular Visualization with PyMOL RMSF values indicate that the spike protein of SARS-CoV-2 RBD is more compatible with binding to the human ACE2 with high flexibility. The results of binding free energies showed a high affinity of SP1-AC2 complex (-52.97 kcal/mol) compared with SP1-CoV2/CD147 (-35.75 kcal/mol). Protein-protein docking was utilized for identifying the hotspot residues in the interface of spike protein with AC2 and CD147. In this paper, we focused our analysis on the initial step of virus infection by comparing the affinity, stability, and specificity of the SARS-CoV-2 spike 1-AC2 and SARS-CoV-2 spike 1-CD147 complexes. CD147, as a biomarker for hyperinflammation, was found to be the functional receptor for SARS-CoV-2 and an additional cell entry route. SARS-CoV-2 invades host cells via interaction of its spike protein with the human angiotensin-converting enzyme 2 as the receptor. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
